As part of a large collaboration with scientists around the world, researchers from the University of California San Diego have developed a new search tool to help researchers better understand the metabolism of microorganisms.
Microbes are key players in virtually all biological and environmental systems, yet limitations in current techniques used to study microbial metabolism make it difficult to decode their interactions and activities.
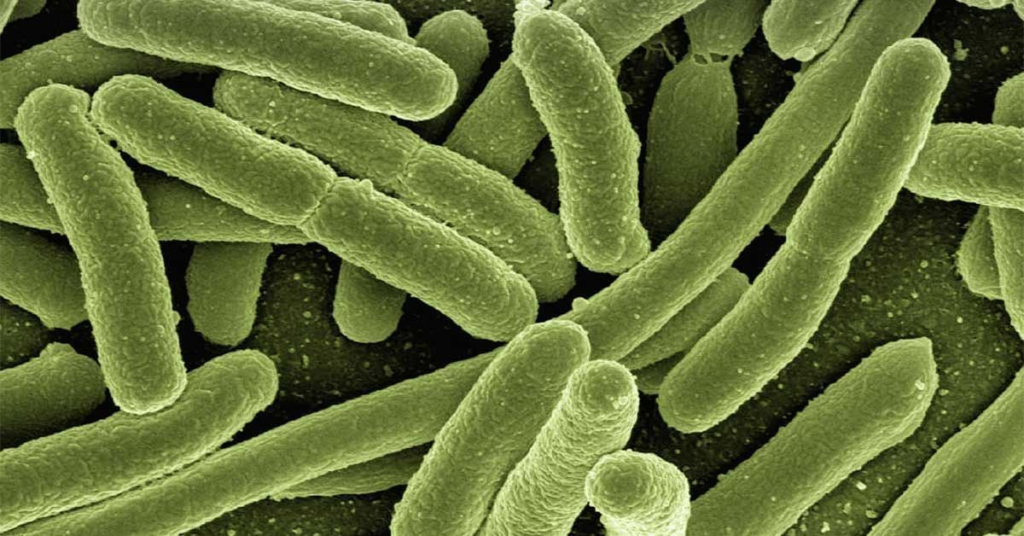
This image shows a microscopic view of E. coli bacteria, a species with an intimate relationship with humans. The new microbeMASST search tool can help research better understand how E. coli and other bacteria affect both human health and the global biosphere.
This image shows a microscopic view of E. coli bacteria, a species with an intimate relationship with humans. The new microbeMASST search tool can help research better understand how E. coli and other bacteria affect both human health and the global biosphere. Image credit: Gerd Altmann/Unsplash
The new research, published in Nature Microbiology, directly addresses these limitations, which could ultimately transform our understanding of both human health and the environment.
“Humans are walking ecosystems in which microbes vastly outnumber us, but we know so little about the metabolites that microbes produce,” said senior study author Pieter Dorrestein, PhD, professor of pharmacology and pediatrics at UC San Diego School of Medicine and professor at Skaggs School of Pharmacy and Pharmaceutical Sciences at UC San Diego.
“This technology allows us to match microbes to the metabolic signatures they produce without any prior knowledge, which represents a major leap forward in our ability to study microorganisms and their intricate relationships with humans and ecosystems.”
The groundbreaking tool, which the scientists call microbeMASST, was developed by scientists at UC San Diego’s Collaborative Microbial Metabolite Center, an NIH-supported initiative that aims to build an internationally-curated repository of microbial metabolomics data to help researchers studying the complex interaction between microbes and humans.
Beneficial microbes play a key role in human health by colonizing certain body areas, including the skin, where they protect us against external pathogens, and the gut, where they contribute to essential functions such as nutrient absorption and regulating the immune system. Disruption of the microbial communities in our body is associated with a wide range of diseases.
“This resource will help us mechanistically interrogate the role of the microbiome in health conditions such as liver disease, inflammatory bowel disease, diabetes, atherosclerosis and others,” added Dorrestein.
Microbes are also at the center of important environmental processes, such as the carbon and nitrogen cycles. When microbial communities are disrupted in these processes, it can become harder for ecosystems to cycle nutrients, leading to a wide range of destructive ecological imbalances.
Because of their crucial role in the environment and their interactions with larger organisms, the metabolism of microbes is a driving force in virtually all aspects of biology. However, the vast metabolic potential of microbial communities is often overlooked in modern experiments, which generally only look at microbial metabolism with a wide lens.
“One of the challenges of studying microbes at the molecular level is that it’s difficult to tell which microbes are producing which molecules unless you already know what you’re looking for,” said first author Simone Zuffa, a postdoctoral researcher working with Dorrestein. “If you think of colonies of microbes as crowded parties with lots of people talking, our current experiments can only record the sound, but we want to figure out a way to unscramble that audio to figure out who is saying what.”
“Humans are walking ecosystems in which microbes vastly outnumber us, but we know so little about the metabolites that microbes produce. This technology allows us to match microbes to the metabolic signatures they produce without prior knowledge, representing a major leap forward in our ability to study microorganisms and their intricate relationships with humans and ecosystems.”Pieter Dorrestein, PhD
To help produce the new search tool, which the researchers have called microbeMASST, researchers from the Collaborative Microbial Metabolite Center at UC San Diego collected more than 100 million data points from 60,000 distinct microbial samples, gathered by scientists from across the world. This database has been meticulously curated from community contributions and metadata curation, and includes microbes from plants, soils, oceans, lakes, fish, terrestrial animals and humans.
By cross-referencing an experimental sample with this massive library of individual microbes, microbeMASST can detect which microbes are present in that sample.
“There’s no existing tool that can do this, and ours can do it in seconds,” added Zuffa.
Because microbeMASST can identify microbes in a sample without any prior knowledge, the researchers are confident that the applications of the technology extend into various fields of biology, such as aquaculture, agriculture, biotechnology, and studying microbial-mediated health conditions.
“We anticipate that microbeMASST will be a transformative resource for the life sciences research community,” said Dorrestein. “Further, the tool will only improve over time as the community gathers more data for the system to reference.”
Source: UCSD

Leave a Reply